We are interested in employing biochemical approaches, in particular, affinity purification of protein complexes coupled with mass spectrometry (AP-MS) technology, for proteomic studies of pluripotent cells including embryonic stem cells (ESCs) and induced pluripotent stem cells (iPSCs).
1. Development of bioSAIP-MS and bioSAIP-chip strategy for studying pluripotency interactome and transcriptional networks in ESCs.
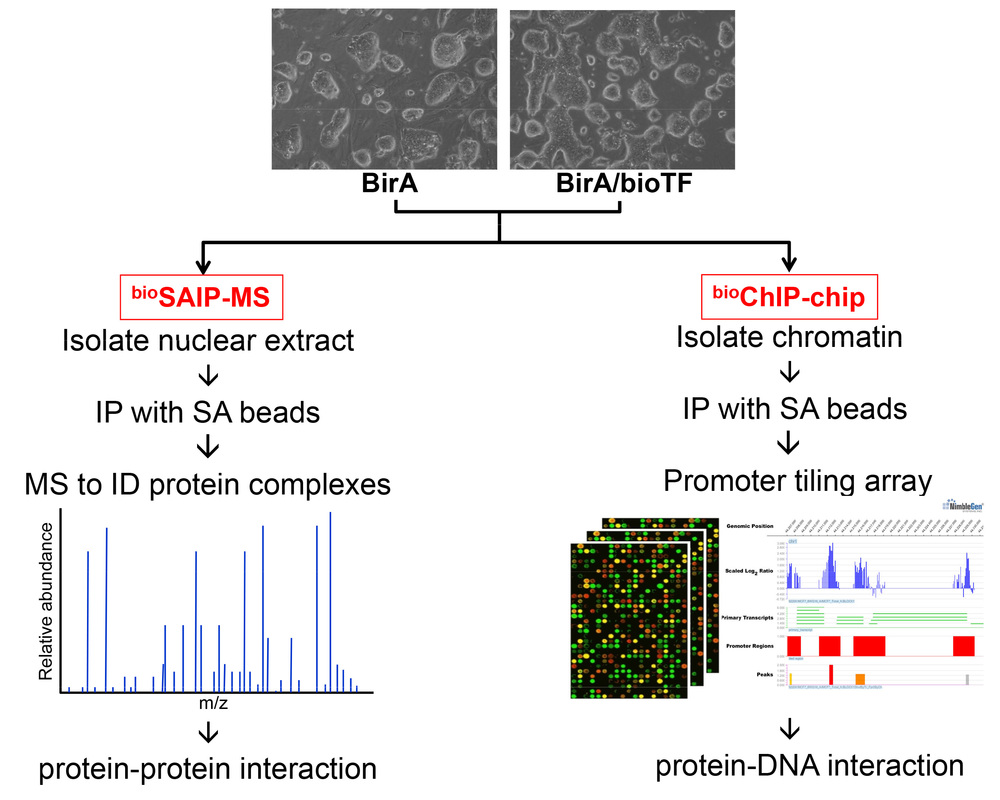
We have tested the utility of in vivo biotinylation of transcription factors in mouse embryonic stem cells (mESCs), and have established an in vivo biotinylation system for BirA-mediated specific biotinylation of critical pluripotency factors in ES cells. We developed and optimized an approach for affinity purification of pluripotency protein complexes involving streptavidin capture of biotinylated proteins (dubbed bioSAIP) and demonstrated the feasibility of in vivo biotinylation for mapping global/chromosomal targets of many different transcription factors (dubbed bioChIP-chip) (Figure 1). Utilizing the technologies we developed, we have constructed a protein interaction network surrounding the pluripotency factor Nanog in mESCs (Wang et al., Nature 2006) and mapped an extended transcriptional network for pluripotency of mESCs (Kim et al. Cell 2008). The network is highly enriched for factors known to be critical in ESC biology and appears to function as a module for pluripotency. Pluripotency is maintained by many transcription factors that form a highly interconnected protein interaction network including the two homeobox proteins Nanog and Oct4, and a battery of associated proteins of known and unknown functions linking to multiple co-repressor pathways.
1. Development of bioSAIP-MS and bioSAIP-chip strategy for studying pluripotency interactome and transcriptional networks in ESCs.
We have tested the utility of in vivo biotinylation of transcription factors in mouse embryonic stem cells (mESCs), and have established an in vivo biotinylation system for BirA-mediated specific biotinylation of critical pluripotency factors in ES cells. We developed and optimized an approach for affinity purification of pluripotency protein complexes involving streptavidin capture of biotinylated proteins (dubbed bioSAIP) and demonstrated the feasibility of in vivo biotinylation for mapping global/chromosomal targets of many different transcription factors (dubbed bioChIP-chip) (Figure 1). Utilizing the technologies we developed, we have constructed a protein interaction network surrounding the pluripotency factor Nanog in mESCs (Wang et al., Nature 2006) and mapped an extended transcriptional network for pluripotency of mESCs (Kim et al. Cell 2008). The network is highly enriched for factors known to be critical in ESC biology and appears to function as a module for pluripotency. Pluripotency is maintained by many transcription factors that form a highly interconnected protein interaction network including the two homeobox proteins Nanog and Oct4, and a battery of associated proteins of known and unknown functions linking to multiple co-repressor pathways.
Figure 1. Strategies for mapping protein-protein and protein-DNA interactions in mESCs. The ESCs expressing BirA alone (as control) and BirA plus biotinylated transcription factors (bioTF) can be used for isolation of protein complexes using streptavidin (SA) immunoprecipitation (IP) coupled with LC-MS/MS (dubbed bioSAIP-MS) and construction of a protein-protein interaction network; meanwhile, the same ESCs can be subjected to in vivo biotinylation-mediated chromatin immunoprecipitation and microarray (dubbed bioChIP-chip) to identify protein-DNA interactions and construct a transcriptional regulatory network.
2. Improving bioSAIP-MS Studies by Establishing bioOct4 and bioNanog ESCs in Null Background.
We have further optimized and improved affinity purification of Oct4 and Nanog protein complexes in mESCs by establishing bioOct4 and bioNanog ESC lines on the genetic null background of respective gene (Figure 2.1 and Figure 2.2). The advantages of these new lines for AP-MS studies are two-fold: 1) there is no endogenous Oct4/Nanog proteins competing with bioOct4/bioNanog for partner proteins; 2) the expression levels of bioOct4/Nanog can be selected and maintained at wild-type levels, which is more abundant than the ectopic expression on the wild-type background (normally kept at 10-20% of the endogenous levels to avoid spurious interactions and maintain the pluripotency status of transgenic ESCs).
By employing these two new ESCs, we have successfully constructed extended Oct4 and Nanog interactomes, which led to our improved understanding of molecular mechanisms underlying stem cell pluripotency and somatic cell reprogramming (Ding et al., Cell Research 2012; Costa, Ding et al., Nature 2013).
We have further optimized and improved affinity purification of Oct4 and Nanog protein complexes in mESCs by establishing bioOct4 and bioNanog ESC lines on the genetic null background of respective gene (Figure 2.1 and Figure 2.2). The advantages of these new lines for AP-MS studies are two-fold: 1) there is no endogenous Oct4/Nanog proteins competing with bioOct4/bioNanog for partner proteins; 2) the expression levels of bioOct4/Nanog can be selected and maintained at wild-type levels, which is more abundant than the ectopic expression on the wild-type background (normally kept at 10-20% of the endogenous levels to avoid spurious interactions and maintain the pluripotency status of transgenic ESCs).
By employing these two new ESCs, we have successfully constructed extended Oct4 and Nanog interactomes, which led to our improved understanding of molecular mechanisms underlying stem cell pluripotency and somatic cell reprogramming (Ding et al., Cell Research 2012; Costa, Ding et al., Nature 2013).
|
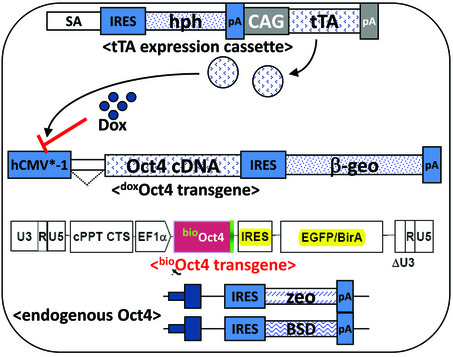
Figure 2.1. Establishment of ZO4B4 ESC line on the Oct4 null background.
This line was established by modifying ZHBTc4 ESCs (Niwa et al., 2000) with lentiviral introduction of bioOct4 transgene. |
|
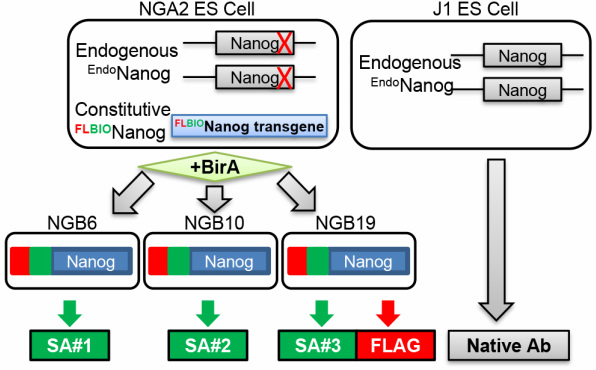
Figure 2.2. Establishment of bioNanog ESC line on the Nanog null background. Schematic depiction of embryonic stem cells expressing NANOG with Flag (FL) and Biotin (BIO) tags (left), and NANOG antibody (Ab)-based affinity purification (right). SA1–SA3, three independent streptavidin pull-down experiments.
|
3. Applying SILAC-based Quantitative MS for Proteomic Studies in Pluripotent Cells.
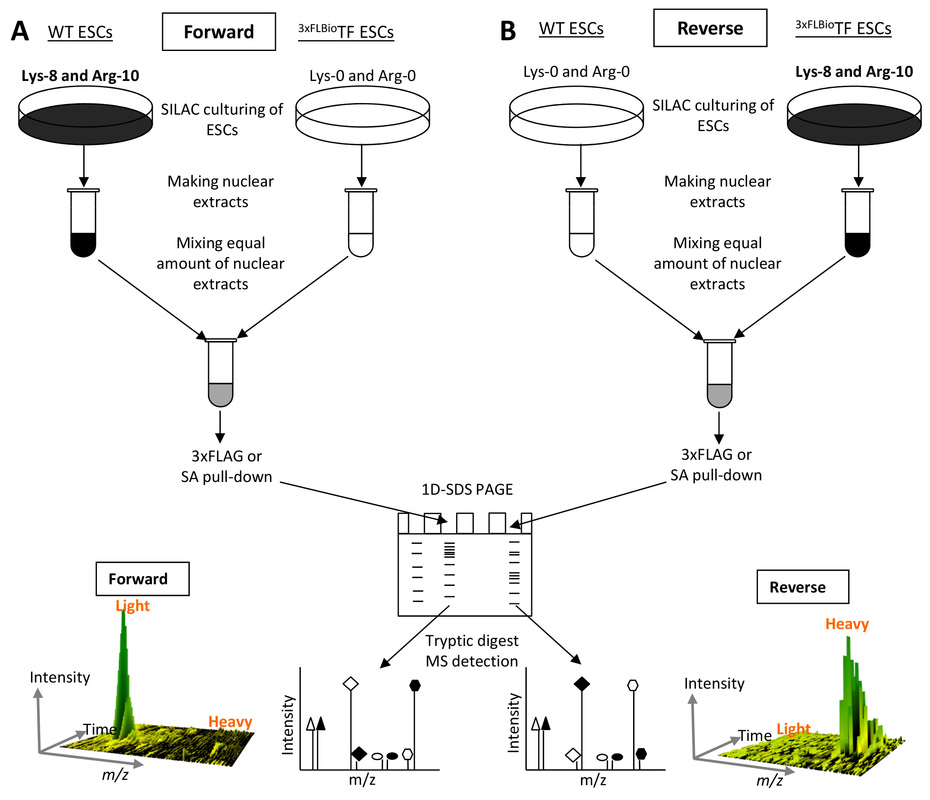
We are currently optimizing the SILAC labeling of ESCs/iPSCs and applying it to AP-MS studies of both mouse and human ESCs/iPSCs (Figure 3). This will greatly enhance our interactome studies.
We are currently optimizing the SILAC labeling of ESCs/iPSCs and applying it to AP-MS studies of both mouse and human ESCs/iPSCs (Figure 3). This will greatly enhance our interactome studies.
Figure 3. SILAC quantitative mass spectrometry for identification of TF-interacting proteins. (A) Forward SILAC strategy; (B) Reverse SILAC strategy.